Application of Functional Genomics to Emerging Model Systems
Life histories vary widely across animals, shaped by ecological factors, developmental constraints, and evolutionary trade‑offs. Our research examines the mechanistic basis of this diversity by investigating how environmental cues interact with physiological and molecular pathways to influence life‑cycle strategies. We focus on animals with complex life cycles—particularly echinoderms, crustaceans, and amphibians—to understand how larval environments affect juvenile and adult phenotypes, including skeletal development, brain growth, immune function, and reproductive output. By integrating endocrine biology, developmental plasticity, and evolutionary theory, we explore how organisms modulate developmental trajectories in response to environmental variability, and how these responses are transmitted across generations. This includes studying transgenerational plasticity, maternal effects, and the evolutionary significance of deferred‑use cell populations. Our goal is to identify general principles that link developmental processes to macroevolutionary patterns, providing a mechanistic foundation for understanding how life histories evolve and respond to rapid environmental change.
Our Work on This Topic
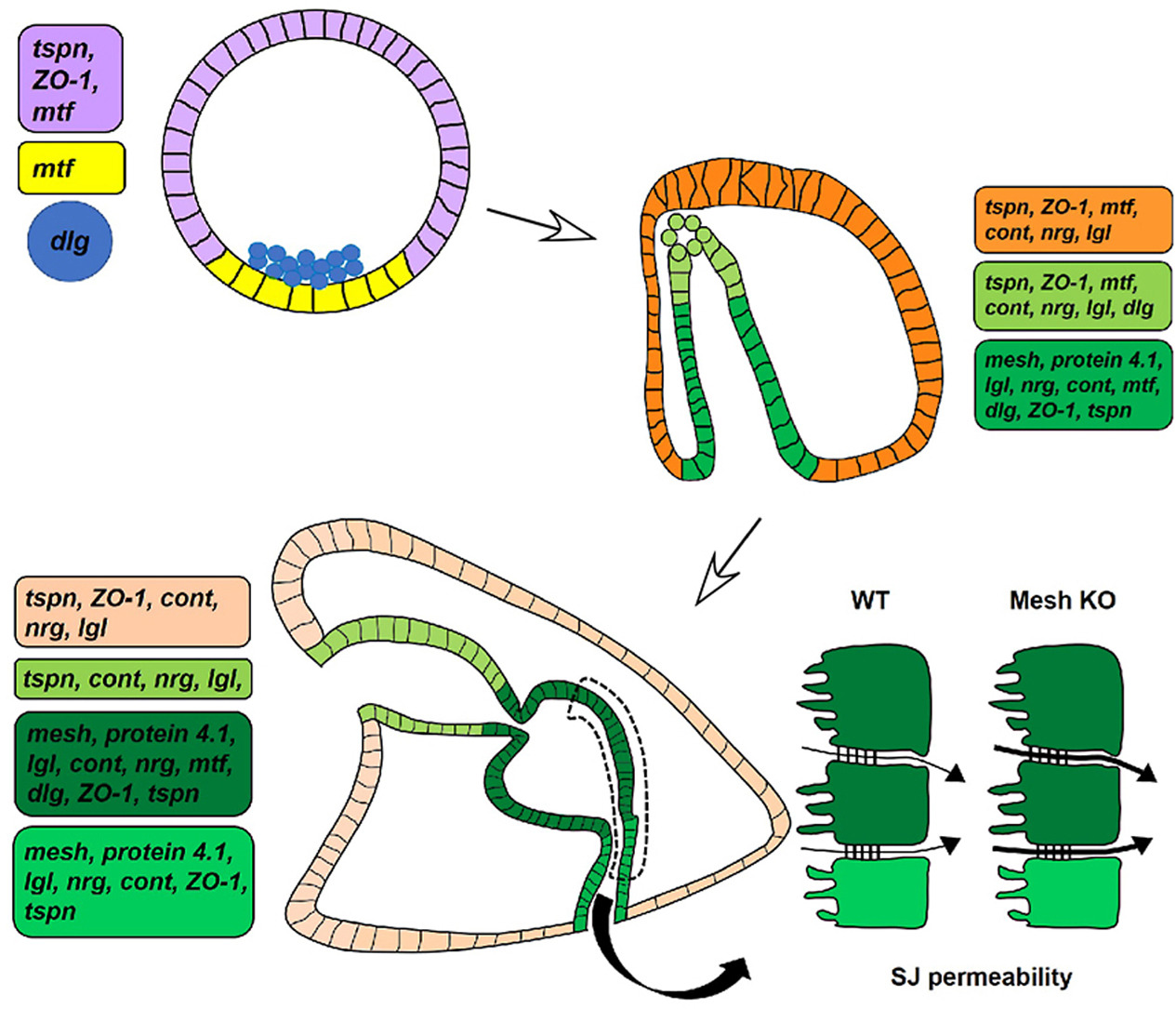
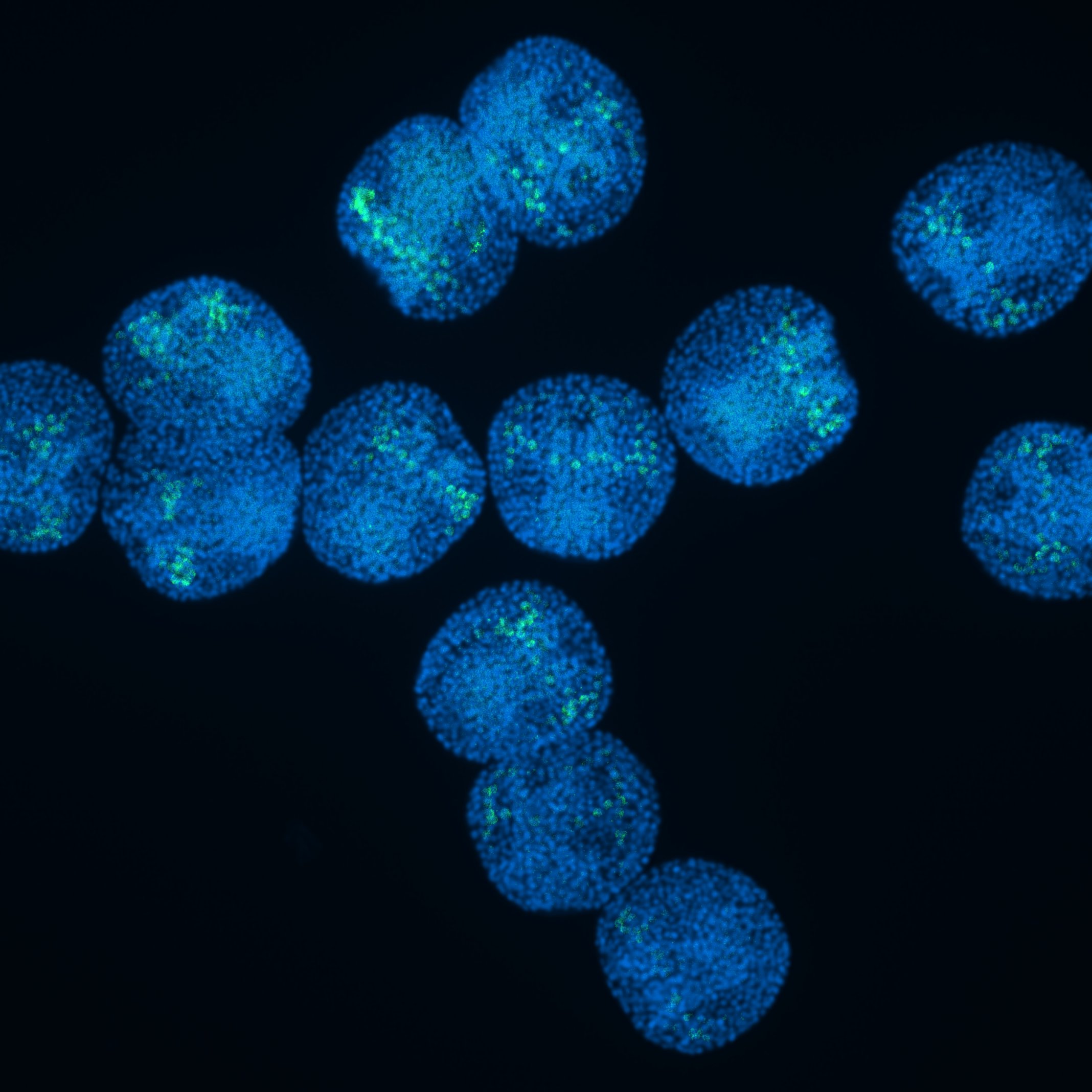
Functional genomics applied to uncover epithelial barrier gene networks in echinoderms.
This chapter presents molecular and cellular techniques—centered on patch‑clamp electrophysiology and cloning—to investigate voltage‑gated calcium channel activity in Trichoplax adhaerens, providing tools to study how this simple placozoan responds to inter‑ and intracellular signaling and offering insight into the evolution of nervous and sensory systems.
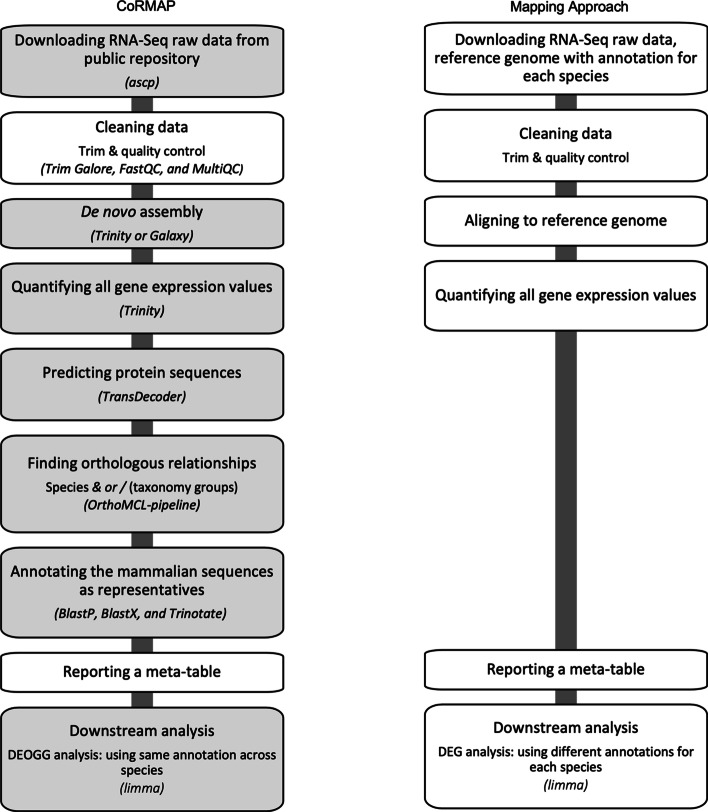
Introduces a versatile pipeline enabling functional genomics in non‑model species.